Research
I am a Lithuanian population geneticist who's interested in the genetic underpinnings of speciation and the evolution of how the genome is organized (in particular sex chromosomes and inversions.) While most of my work is theory driven, I think one of the most exciting aspects of the age of genomic data is that we can at last use real data to parameterize and test our models. Currently, I am studying broad patterns of speciation and introgression across groups in Daniel Matute's lab at UNC Chapel Hill as a postdoctoral researcher, but I will be starting my own group at the University of Connecticut in Spring 2024. I am looking for graduate students and postdocs, so if my work interests you and you would like to join, feel free to email adagilis(at)uconn.edu
Speciation
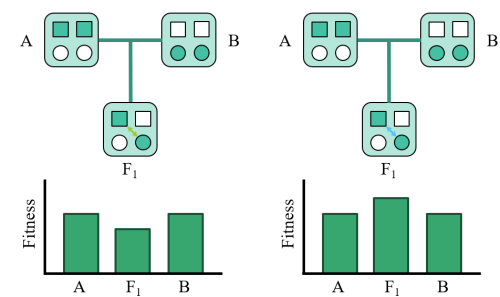
What makes two perfectly adapted populations stop being able to interbreed? I tend to approach this question by looking at the interactions between genes, and asking how the landscape of these interactions impacts the fate of evolving populations. I think this question is particularly exciting, since many parallels outside of biology exist (e.g. languages, branches of software, legal systems, etc.). Is the break-down of hybrid fitness observed in natural populations a general outcome for any two systems that diverge, or are some properties unique to biology?
Sex Chromosomes
While I'm intersted in the evolution of genome organization broadly, I think one of the most dynamic elements of many genomes is the sex chromosomes. Much of our theory and knowledge of sex chromosomes has been based on the rather old and slow to change systems in mammals and birds. However, research in the last few decades has demonstrated that these taxa represent a drastically different view of sex chromosomes than do many fish, amphibians and insects. However, time and time again studies find that these are unique elements when it comes to their role in speciation, introgression, development and many other biological processes. What are the drivers of the evolution of sex chromosomes and their maintenance? What determines the eventual degredation or turnover of these chromosomes?
Genomic inferences
One of the joys of being a population geneticist in our times is that many parameters that simply had to be assumed are now beginning to be known. On the basic end this means we know the number of genes, mutation rates of those genes, and something about the effective population size of different species. On the more complex end of the scale, we are beginning to truly measure epistatic interactions among vast numbers of mutations, computational methods allow us to get a good idea of the demographic changes in and between species and the selective forces that shape our genome are being measured. This presents a great opportunity for population geneticists to both participate in the analysis of genomic datasets to help infer these previously unknowable parameters, but also to test hypotheses that previously were intractible. Do genes that interact, for instance, end up more closely linked in the genome? Are the epistatic interactions observed in real data sufficient to explain rates of speciation? Is introgression common only between recent species, or is it something that happens even between highly diverged groups? Below is some introgression data (just plants) from a recent meta-analysis the Matute Lab and I performed. There is a large degree of variation in the amount of genome showing evidence for introgression (roughly proportional to Patterson's D) and genetic distance (here measured as Jukes-Cantor), and some of this variation seems to be taxon specific (you can toggle which orders are displayed by clicking them in the legend).
Medical Mycology
I have recently participated in an NIH T32 training grant for Medical Mycology and Molecular Pathogenesis. This program allowed me to apply many of the genomic techniques I have learned to something more practical - examining the evolutionary history of pathogenic fungi. These fungi are a group of organisms that have historically posed danger primarily to the global south and as a result have received very little attention in science. As part of the training program, I am helping the Matute lab look into species boundaries among Histoplasma species as well as examining the evolutionary relationships between different strains, the evolution of structural variation in these organisms and examining how reference bias impacts evolutionary inference in these (often highly structurally variable) species.
Join the lab!
If you are an undergraduate student at UConn who is excited to develop new population genetics approaches to thinking about speciation and hybridization, I would love to hear from you!
If you are looking for a graduate program that will let you learn how to develop theory, methods and do so in a vibrant and supportive department - I am currently looking for students.
Finally, if you are a graduate student who is excited to pick up some population genetics skills in an environment that encourages you (and has the funding to) let you develop and explore your own system, think about a postdoctoral position in the lab.
Reach out at adagilis(at)uconn.edu
Publications
Up to date list on Google Scholar
Publications
2023
- Dagilis, A.J., Matute D.R., The fitness of an introgressing haplotype changes over the course of divergence and depends on its size and genomic location. PLoS Biology
2022
- Dagilis, A. J., Peede D., Coughlan J. M., Jofre G. I., D'Agostino E. R. R., Mavengere H., Tate A. D., Matute D. R. (2022). A need for standardized reporting of introgression: Insights from studies across eukaryotes. Evolution Letters
- Dagilis, A. J., Sardell J. M., Josephson M. P., Su Y., Kirkpatrick M., Peichel C.L. (2022). Searching for signatures of antagonistic selection on stickleback sex chromosomes. Philosophical Transactions of the Royal Society B, 377(1856)
- Dagilis, A. J. (2022). What inversion lengths can tell us about their evolution. Molecular Ecology, 31(13) - News and Views
2020
- Dagilis, A. J., & Matute, D. R. (2020). Incompatibilities between emerging species. Science, 368(6492), 710-711. - News and Views
2019
- Dagilis, A. J., Kirkpatrick, M., & Bolnick, D. I. (2019). The evolution of hybrid fitness during speciation. PLoS genetics, 15(5). pdf
2018
- Sardell, J.M., Cheng C., Dagilis, A. J., Ishikawa A., Kitano J., Peichel C.L., Kirkpatrick M, (2018). Sex differences in recombination in sticklebacks. G3: Genes, Genomes, Genetics, 8(6).
- Kuzmin E., VanderSluis B.,... Dagilis, A. J. ..., Boone C., Myers C. (2018). Systematic analysis of complex genetic interactions. Science 360(6386).
2016
- Dagilis, A. J., Kirkpatrick, M. (2016). Prezygotic isolation, mating preferences, and the evolution of chromosomal inversions. Evolution, 70(7). url. pdf
2012
- Livingstone K., Olofsson P., Cochran G., Dagilis, A. J., MacPherson K., Seitz K. (2012) A stochastic model for the development of Bateson-Dobzhansky-Muller incompatibilities that incorporates the structure of protein interaction networks. Mathematical biosciences 238(1).
Resume
Dagilis lab arriving at UConn in Spring 2024!
Currently a postdoc at UNC Chapel Hill in Daniel Matute's Lab. CV
Summary
Andrius J. Dagilis
Postdoctoral fellow
- Matute Lab
- University of North Carolina, Chapel Hill
- Genome Sciences Building 3101
- adagilis@uconn.edu
Education
PhD; Ecology, Evolution and Behavior
2013 - 2019
University of Texas, Austin, TX
Advised by Mark Kirkpatrick, worked on speciation, sex chromosomes, and inversions.
Bachelor of Sciences; Biology
2009 - 2013
Trinity University, San Antonio, TX
Worked in Kevin Livingstone and Peter Olofsson's labs, developed a long standing interest in speciation and biomathematics.
Skills and Interests
General competence in
- Mathematical modeling
- Population genomics
- Shell scripting
- Analysis and plotting in R
Outside of work
- Running
- Beer brewing
- Any volunteer work that includes removing invasive species
- Showing my lack of competence at field work by helping out
